 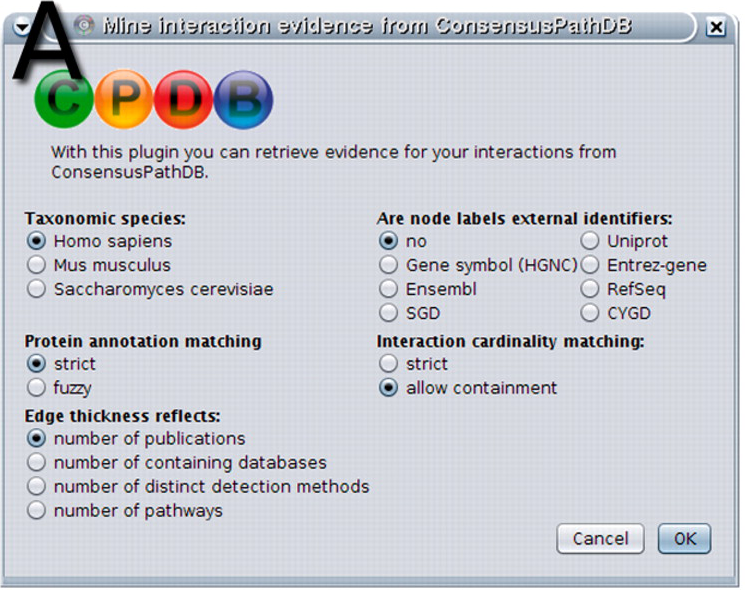
k Kernel width (accurate method only), default: determined by program a Algorithm (fixed_bandwidth | variable_bandwidth | adaptive_partitioning), default: adaptive_partitioning See or Download PDF and Supplemental Documents for detailed usage instructions. Please set a 'JAVA_HOME' environment variable pointing to your JDK and use the launch_aracne scripts in the distribution to start the application Documentation and Support Java Graphic User Interface (GUI) for loading adjacency matrices and drawing network diagrams using a built-in Cytoscape plugin. PE32 executable for MS Windows (console) Intel 80386 32-bitĮLF 64-bit LSB executable, AMD x86-64, version 1 (SYSV), for GNU/Linux 2.6.9, dynamically linked (uses shared libs), for GNU/Linux 2.6.9, not stripped Please copy this file to the same directory as the binary (This can be ignored if the entire source distribution is downloaded) This file is used by the native aracne2 binaries compiled from C++ source to provide ARACNe2 usage summary. Application DownloadĪvailable formats for ARACNe2 include the following. Click here to download the Cytoscape app. Cytoscape AppĪRACNe is now available as a Cytoscape app that is integrated in the Cyni Toolbox panel. GPU-ARACNE provides a parallelized implementation of ARACNe for GPU architectures. ARACNe-APĪRACNe-AP implements adaptive partitioning to improve the scalability of the original implementation. Application to the deconvolution of genetic networks in human B cells demonstrates ARACNe’s ability to infer validated transcriptional targets of the c-MYC proto-oncogene. On synthetic datasets ARACNe achieves extremely low error rates and significantly outperforms established methods, such as Relevance Networks and Bayesian Networks. This method uses an information theoretic approach to eliminate the vast majority of indirect interactions typically inferred by pairwise analysis. Installing it provides many useful additional layouts, especially with large networks with many nodes and connections.ARACNe (Algorithm for the Reconstruction of Accurate Cellular Networks) is a novel algorithm, using microarray expression profiles, that is specifically designed to scale up to the complexity of regulatory networks in mammalian cells, yet general enough to address a wider range of network deconvolution problems. I also recommend the free add-on yFiles Layout Algorithms from the Cytoscape app store. 
The CSV can then be imported into Cytoscape where it generates the corresponding network diagrams (nodes, edges, attributes, labels, etc). The raw netstat output would need only a little cleaning up to convert it into a CSV. Cytoscape core distribution provides a basic set of features for data integration, analysis, and visualization. Although Cytoscape was originally designed for biological research, now it is a general platform for complex network analysis and visualization. Although it originates from the biological space, it is totally usable in any other graph and/or relationship context.Ĭytoscape is an open source software platform for visualizing molecular interaction networks and biological pathways and integrating these networks with annotations, gene expression profiles and other state data. I had to solve a similiar problem recently (DB object dependencies) and after a lot of research I ended up using Cytoscape, an open source graph visualisation tool. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
